In silico functional and tumor suppressor role of hypothetical protein PCNXL2 with regulation of the Notch signaling pathway - RSC Advances (RSC Publishing) DOI:10.1039/C8RA00589C

Discover the new Expasy.org, the Swiss Bioinformatics Resource Portal | SIB Swiss Institute of Bioinformatics

Physiochemical Characterization and Domain Annotation of ORF1ab Polyprotein of Novel Corona Virus 19 | SpringerLink

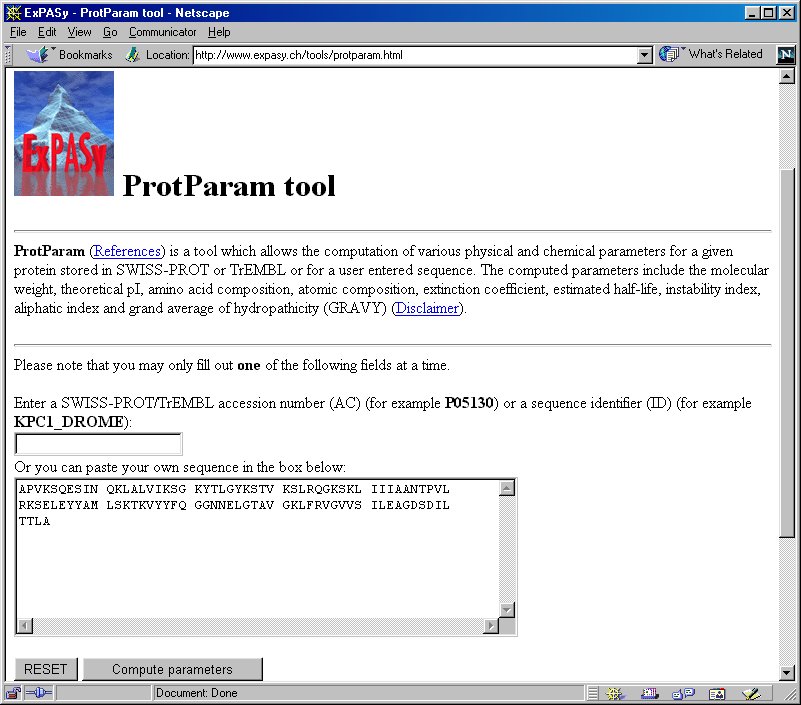
SOLVED: Use EXPASY Protparam tool to calculate the molecular weight, number of amino acids, and the theoretical pI, extinction coefficient under fully reducing conditions, and expected absorbance for 1 mg/ml protein. Use

SOLVED: Use EXPASY Protparam tool to calculate the molecular weight, number of amino acids, and the theoretical pI, extinction coefficient under fully reducing conditions, and expected absorbance for 1 mg/ml protein. Use